Research
Deciphering the Genomic Rosetta Stone

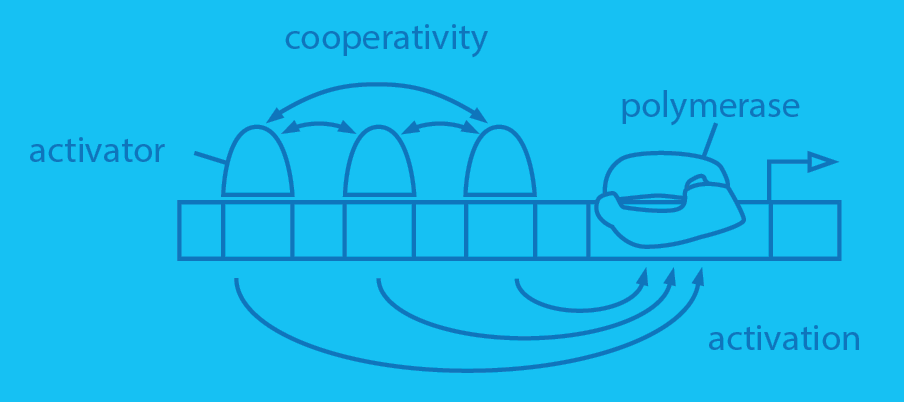
Most people surveyed would say that Escherichia coli is probably biology’s best understood organism. And yet, out of its more than 4500 different genes, we know nothing about how roughly half of those genes are regulated. Literally nothing – no transcription factor binding sites, no transcription factors identified that regulate those genes. Work in our laboratory is using a combination of mutagenesis, cell-sorting, deep sequencing, mass spectrometry and models from information theory and statistical physics to uncover the regulatory architectures of some of these uncharacterized genes. Once we have these regulatory architectures in hand, we then turn to allied efforts in the laboratory aimed at using statistical mechanical models of gene regulation in conjunction with precision measurements to predict and measure the full input-output behavior of these genes. Further, we are also engaged in using these approaches in the evolutionary setting to explore how transcription factors and their binding sites co-evolve.

Tom

Kian

Rosalind
Specificity Costs
“Napoleon is in equilibrium”. These words were fired off at a speaker who had used ideas from equilibrium statistical mechanics to evaluate some process in molecular biophysics. Though equilibrium ideas certainly work in many contexts that at first blush seem inappropriate, indeed, there are many biological processes which are highly dependent upon the fact that the system is out of equilibrium. Work in our group in this area aims to explore how processes such as DNA replication have such amazingly high fidelity in comparison with what is permitted by thermodynamic equilibrium alone. A second exciting set of questions center on how energy invested in biological processes permits reactions to occur in a sequential fashion that is forbidden by equilibrium considerations.

Gabe

Sara
Active Matter


Heun Jin

Soichi

Ana